IGF-1LR3 1mg (3ml)
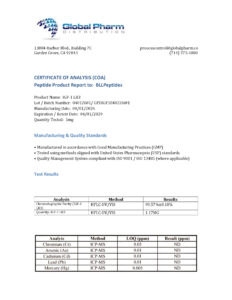
IGF-1 LR3 is a research-grade long-acting Insulin-like Growth Factor 1 analog studied for potent anabolic effects on muscle protein synthesis, cellular proliferation, and glucose uptake via IGF-1 receptor signaling. Researchers investigating growth factor biology and anabolic physiology rely on pharmaceutical-grade purity for accurate data. Available at BLL Peptides — USA-made, rigorously tested. ✅ COA tested…
Description
IGF-1 LR3: Complete Research Guide – Long Arginine Insulin-Like Growth Factor Mechanisms, Anabolic Research, and Cell Proliferation Applications
Last updated: March 2026
Executive Summary
IGF-1 LR3 (Long Arginine 3 Insulin-Like Growth Factor 1) is a synthetic analog of human insulin-like growth factor 1 (IGF-1) that has been engineered for dramatically enhanced biological potency and extended activity. The analog consists of 83 amino acids compared to the 70-amino acid native IGF-1, incorporating a 13-amino acid N-terminal extension peptide and a critical glutamic acid-to-arginine substitution at position 3 of the native sequence (Glu3 to Arg3). These modifications reduce the binding affinity for IGF-binding proteins (IGFBPs) by approximately 100-fold, resulting in a substantially greater proportion of free, bioactive peptide in circulation [1, 2].
Native IGF-1 circulates predominantly in ternary complexes with IGFBP-3 and the acid-labile subunit (ALS), which limits its bioavailability and confers a plasma half-life of only 12-15 minutes for free IGF-1. IGF-1 LR3, by contrast, exhibits a functional half-life of approximately 20-30 hours due to its dramatically reduced sequestration by IGFBPs. This extended bioavailability translates to far greater activation of the IGF-1 receptor (IGF-1R) and downstream PI3K/Akt/mTOR signaling cascades that govern protein synthesis, cell proliferation, and anti-apoptotic responses [3, 4].
With a molecular weight of approximately 9,111 Da and CAS number 946870-92-4, IGF-1 LR3 has become an indispensable tool in cell biology research, serving as a potent cell culture media supplement that promotes cell growth and survival. In the broader research landscape, it has been studied extensively in contexts ranging from skeletal muscle hypertrophy and satellite cell activation to cancer biology and metabolic signaling. This guide provides a comprehensive, evidence-based examination of IGF-1 LR3's molecular structure, mechanisms of action, published research findings, safety profile, and applications in contemporary scientific investigation.
Interactive Molecular Structure
The following interactive 3D visualization renders the IGF-1 LR3 peptide backbone in an alpha-helical representation. The structure highlights the engineered N-terminal extension (residues 1-13, purple) that distinguishes it from native IGF-1, the critical Arg16 mutation site (position 3 of the native sequence, orange), and the native IGF-1 core domains (residues 14-83). IGF-1 LR3 contains three disulfide bonds that stabilize its tertiary structure.
Legend: The interactive visualization above depicts the 83-residue backbone of IGF-1 LR3. The N-terminal extension (residues 1-13, purple) is the engineered addition absent from native IGF-1. The orange-highlighted Arg16 (R*16) marks the critical Glu-to-Arg substitution at position 3 of the native IGF-1 sequence that reduces IGFBP binding by approximately 100-fold. Gold dashed lines indicate the three disulfide bonds (Cys-Cys) that stabilize the protein's tertiary fold. The C domain loop region (residues 43-51) is shown with wider, looser geometry reflecting its flexible structure. Drag to rotate; scroll to zoom.
Table of Contents
- Introduction and Discovery History
- Molecular Structure and Chemistry
- Mechanism of Action
- Scientific Research Review
- Comparison with Related Growth Factors
- Safety Profile and Pharmacology
- Research Applications
- References
- Disclaimer
Introduction and Discovery History
The Discovery of IGF-1
The story of insulin-like growth factor 1 begins in the late 1950s, when researchers sought to understand the mechanism by which growth hormone (GH) stimulated somatic growth. In 1957, Salmon and Daughaday proposed the "somatomedin hypothesis," observing that growth hormone did not directly stimulate sulfate incorporation into cartilage but rather acted through a serum factor they termed "sulfation factor" [5]. This intermediary was later renamed somatomedin C and ultimately identified as insulin-like growth factor 1 (IGF-1).
The complete amino acid sequence of human IGF-1 was determined in 1978 by Rinderknecht and Humbel, revealing a 70-amino acid single-chain polypeptide with approximately 50% sequence homology to proinsulin [6]. This structural similarity to insulin explained the earlier observation that serum contained factors with "insulin-like" activity that could not be suppressed by anti-insulin antibodies — the so-called "non-suppressible insulin-like activity" (NSILA). IGF-1 was found to be primarily produced in the liver under growth hormone stimulation, though paracrine and autocrine production occurs in virtually all tissues.
The GH-IGF-1 axis was subsequently established as the principal endocrine pathway governing postnatal longitudinal growth, with IGF-1 mediating the majority of growth hormone's anabolic and mitogenic effects. Clinical studies in patients with growth hormone insensitivity (Laron syndrome), who have normal or elevated GH but extremely low IGF-1 levels, confirmed the essential role of IGF-1 in human growth and development [7].
The IGF Binding Protein Problem
A critical challenge in IGF-1 biology emerged with the discovery that more than 99% of circulating IGF-1 is bound to a family of six high-affinity IGF binding proteins (IGFBP-1 through IGFBP-6). The predominant circulating complex consists of IGF-1 bound to IGFBP-3 and the acid-labile subunit (ALS), forming a approximately 150 kDa ternary complex. This complex serves as a reservoir that extends the half-life of total IGF-1 to approximately 12-16 hours, but the free (unbound) IGF-1 that can actually activate the IGF-1 receptor has a half-life of only 12-15 minutes [8].
The IGFBPs function as both reservoirs and regulators. IGFBP-3, which carries approximately 75-80% of circulating IGF-1, modulates IGF-1 bioavailability through controlled release. However, this tight binding also severely limits the biological potency of exogenous IGF-1 administered in research settings. When native IGF-1 is added to experimental systems, the majority is rapidly sequestered by endogenous IGFBPs, dramatically reducing its effective concentration at the receptor [9].
This limitation motivated the development of IGF-1 analogs with reduced IGFBP binding affinity, culminating in the creation of IGF-1 LR3.
Engineering IGF-1 LR3
IGF-1 LR3 was developed through structure-function studies of the IGF-1/IGFBP interaction interface. Crystallographic and mutagenesis research identified glutamic acid at position 3 (Glu3) of native IGF-1 as a critical contact residue for IGFBP binding. The Glu3 residue, located in the N-terminal region of the B domain, makes direct electrostatic contacts with multiple IGFBPs [10].
Two key modifications were introduced to create the LR3 variant:
N-terminal extension: A 13-amino acid peptide (MFPAMPLSSLFVN) was appended to the N-terminus of native IGF-1. This extension, derived from methionyl porcine growth hormone, sterically disrupts the IGFBP binding interface by altering the spatial presentation of the N-terminal region [2].
Arg3 substitution: Glutamic acid at position 3 of the native IGF-1 sequence (corresponding to position 16 in the LR3 numbering) was replaced with arginine. This charge reversal — from negative (Glu) to positive (Arg) — abolishes a critical electrostatic interaction with the IGFBPs. The combination of charge reversal and steric effects from the N-terminal extension reduces IGFBP binding affinity by approximately 100-fold compared to native IGF-1 [1, 2].
Importantly, these modifications do not significantly impair binding to the IGF-1 receptor (IGF-1R), which recognizes a different epitope on the IGF-1 molecule. The receptor binding site involves residues in the B and A domains that are distant from the N-terminal modification, allowing IGF-1 LR3 to retain full agonist activity at IGF-1R while evading IGFBP sequestration [11].
The result is a peptide with dramatically enhanced and prolonged biological activity: IGF-1 LR3 remains in the free, bioactive form for approximately 20-30 hours, compared to the approximately 12-15 minutes for free native IGF-1.
Molecular Structure and Chemistry
Primary Structure
IGF-1 LR3 is an 83-amino acid single-chain polypeptide with the following complete sequence:
Full sequence (83 residues):
Met-Phe-Pro-Ala-Met-Pro-Leu-Ser-Ser-Leu-Phe-Val-Asn-Gly-Pro-Arg-Thr-Leu-Cys-Gly-Ala-Glu-Leu-Val-Asp-Ala-Leu-Gln-Phe-Val-Cys-Gly-Asp-Arg-Gly-Phe-Tyr-Phe-Asn-Lys-Pro-Thr-Gly-Tyr-Gly-Ser-Ser-Ser-Arg-Arg-Ala-Pro-Gln-Thr-Gly-Ile-Val-Asp-Glu-Cys-Cys-Phe-Arg-Ser-Cys-Asp-Leu-Arg-Arg-Leu-Glu-Met-Tyr-Cys-Ala-Pro-Leu-Lys-Pro-Ala-Lys-Ser-Ala
Where Arg (bold, position 16) represents the critical Glu-to-Arg substitution.
Structural Domains
The IGF-1 LR3 molecule can be divided into five structural regions:
| Domain | Residues (LR3 numbering) | Native IGF-1 equivalent | Description |
|---|---|---|---|
| N-terminal extension | 1-13 | Not present | Engineered extension from methionyl porcine GH |
| B domain | 14-42 | 1-29 | Alpha-helical region, contains receptor binding determinants |
| C domain | 43-51 | 30-38 | Flexible loop connecting B and A domains |
| A domain | 52-75 | 39-62 | Alpha-helical region, disulfide-bonded core |
| D domain | 76-83 | 63-70 | Short C-terminal extension |
Disulfide Bond Architecture
IGF-1 LR3 contains six cysteine residues forming three intramolecular disulfide bonds that are essential for maintaining the bioactive three-dimensional structure:
- Cys19-Cys61 (LR3 numbering; Cys6-Cys48 in native IGF-1): Connects the B domain to the A domain
- Cys31-Cys65 (Cys18-Cys52 in native): Links within the core, stabilizing the hydrophobic interior
- Cys61-Cys74 (Cys48-Cys61 in native): An intra-A domain bond critical for helix stabilization
This disulfide bond pattern is analogous to that found in insulin, consistent with the evolutionary relationship between the insulin and IGF families. Reduction of these disulfide bonds results in complete loss of biological activity, as the properly folded tertiary structure is required for IGF-1R recognition [12].
Physicochemical Properties
| Property | Value |
|---|---|
| Molecular formula | C400H625N111O115S9 |
| Molecular weight | Approximately 9,111 Da |
| CAS number | 946870-92-4 |
| Isoelectric point (pI) | Approximately 8.4 |
| Number of amino acids | 83 |
| Disulfide bonds | 3 |
| Appearance | White lyophilized powder |
| Solubility | Soluble in water, 10 mM HCl, or acetic acid |
| Storage stability | -20C lyophilized (long-term); 4C in solution (short-term) |
Comparison with Native IGF-1 Structure
The three-dimensional structure of native IGF-1 has been determined by both NMR spectroscopy and X-ray crystallography. The molecule adopts a compact globular fold stabilized by its three disulfide bonds, with the B and A domain helices arranged in a configuration reminiscent of insulin. The C domain forms a flexible loop connecting these two helical segments, while the D domain extends as a short, relatively disordered C-terminal tail [12].
In IGF-1 LR3, the core structure (residues 14-83) retains the same overall fold as native IGF-1. The N-terminal extension (residues 1-13) projects outward from the B domain helix, creating steric bulk in the region where IGFBPs normally dock. This engineered extension does not fold into a stable secondary structure but rather exists as a flexible appendage that physically blocks IGFBP access to the N-terminal binding determinants [2, 10].
Mechanism of Action
IGF-1 Receptor Activation
IGF-1 LR3 exerts its biological effects primarily through the IGF-1 receptor (IGF-1R), a transmembrane receptor tyrosine kinase that shares approximately 60% sequence homology with the insulin receptor [13]. The IGF-1R is a heterotetrameric receptor composed of two extracellular alpha subunits and two transmembrane beta subunits linked by disulfide bonds.
Upon binding of IGF-1 LR3 to the alpha subunits of IGF-1R, the receptor undergoes a conformational change that activates the intrinsic tyrosine kinase activity of the beta subunit intracellular domains. This results in auto-phosphorylation of multiple tyrosine residues within the beta subunit, creating docking sites for downstream signaling adaptors. The two major signaling cascades activated by IGF-1R are:
- PI3K/Akt/mTOR pathway: Mediating metabolic, anti-apoptotic, and protein synthesis effects
- Ras/Raf/MEK/ERK (MAPK) pathway: Mediating mitogenic and differentiation signals
Because IGF-1 LR3 retains full affinity for IGF-1R while evading IGFBP sequestration, it achieves far greater receptor occupancy per mole of administered peptide than native IGF-1. This translates to proportionally greater activation of downstream signaling cascades [3, 4].
The PI3K/Akt/mTOR Signaling Cascade
The PI3K/Akt/mTOR pathway is the principal mediator of IGF-1 LR3's anabolic and survival effects. The signaling sequence proceeds as follows:
Step 1 — IRS recruitment: Activated IGF-1R phosphorylates insulin receptor substrate (IRS) proteins, primarily IRS-1 and IRS-2, on specific tyrosine residues. These phosphorylated IRS proteins serve as scaffolds for downstream signaling complex assembly [14].
Step 2 — PI3K activation: Phosphorylated IRS recruits and activates phosphoinositide 3-kinase (PI3K), a lipid kinase that converts phosphatidylinositol-4,5-bisphosphate (PIP2) to phosphatidylinositol-3,4,5-trisphosphate (PIP3) at the inner leaflet of the plasma membrane.
Step 3 — Akt activation: PIP3 recruits Akt (protein kinase B) to the membrane via its pleckstrin homology (PH) domain, where Akt is phosphorylated and activated by PDK1 (at Thr308) and mTORC2 (at Ser473).
Step 4 — mTORC1 activation: Activated Akt phosphorylates and inactivates TSC2 (tuberin), releasing its inhibition of Rheb (Ras homolog enriched in brain). Active Rheb then directly stimulates mTORC1 (mechanistic target of rapamycin complex 1).
Step 5 — Protein synthesis: mTORC1 phosphorylates two key effectors: p70S6 kinase (S6K1), which enhances ribosomal protein synthesis, and 4E-BP1, releasing the translation initiation factor eIF4E to promote cap-dependent mRNA translation [15].
This cascade is the molecular basis for IGF-1 LR3's powerful stimulation of protein synthesis and cell growth. In skeletal muscle, mTORC1 activation by IGF-1 signaling is one of the primary drivers of muscle protein synthesis and hypertrophy [16].
Anti-Apoptotic Mechanisms
IGF-1 LR3 promotes cell survival through multiple Akt-dependent anti-apoptotic mechanisms:
BAD phosphorylation: Akt directly phosphorylates the pro-apoptotic protein BAD at Ser136, causing BAD to dissociate from the anti-apoptotic protein Bcl-2 and associate instead with 14-3-3 scaffolding proteins. This liberates Bcl-2 and Bcl-xL to protect mitochondrial membrane integrity and prevent cytochrome c release [14].
Caspase-9 inactivation: Akt phosphorylates procaspase-9, preventing its activation and thereby blocking the intrinsic (mitochondrial) apoptotic pathway.
FOXO transcription factor inhibition: Akt phosphorylates FOXO family transcription factors (FOXO1, FOXO3a), causing their nuclear exclusion and preventing transcription of pro-apoptotic genes including FasL, Bim, and TRAIL [17].
NF-kappaB activation: Through phosphorylation of IKK (IkappaB kinase), Akt can promote NF-kappaB nuclear translocation and expression of survival genes.
These converging anti-apoptotic mechanisms make IGF-1 LR3 a potent survival factor for diverse cell types, which is a key reason for its widespread use as a cell culture supplement to maintain cell viability and prevent serum-starvation-induced cell death.
Ras/MAPK Proliferative Signaling
In parallel with the PI3K/Akt pathway, IGF-1R activation by IGF-1 LR3 stimulates the Ras/Raf/MEK/ERK mitogen-activated protein kinase (MAPK) cascade:
Ras activation: Phosphorylated IRS and Shc adaptor proteins recruit the Grb2-SOS complex to the membrane, where SOS functions as a guanine nucleotide exchange factor (GEF) to activate Ras by promoting GDP-to-GTP exchange [18].
Kinase cascade: Active Ras recruits and activates Raf (MAPKKK), which phosphorylates and activates MEK1/2 (MAPKK), which in turn phosphorylates and activates ERK1/2 (MAPK). ERK1/2 translocates to the nucleus where it phosphorylates transcription factors including Elk-1, c-Fos, and c-Myc.
Cell cycle progression: ERK-dependent transcription drives expression of cyclin D1 and other cell cycle regulators, promoting G1-to-S phase transition and cell proliferation.
The relative activation of PI3K/Akt versus MAPK pathways determines the balance between pro-survival/anabolic responses and proliferative responses, and varies by cell type and context [19].
Enhanced Bioavailability: The IGFBP Evasion Mechanism
The defining pharmacological advantage of IGF-1 LR3 is its dramatically reduced binding to IGF-binding proteins. In the native IGF-1 system, six IGFBPs (IGFBP-1 through IGFBP-6) compete with the IGF-1R for IGF-1 binding, effectively buffering free IGF-1 concentration to very low levels (less than 1% of total IGF-1) [8].
The binding affinities of native IGF-1 for various IGFBPs range from approximately 0.1 to 10 nM (Kd), which are comparable to or greater than its affinity for IGF-1R (approximately 0.2-1 nM Kd). This means that in any biological system containing IGFBPs, the majority of native IGF-1 will be sequestered and unavailable for receptor activation.
IGF-1 LR3's approximately 100-fold reduction in IGFBP affinity (Kd values of approximately 10-1000 nM for various IGFBPs) while retaining near-native IGF-1R affinity means that in the presence of IGFBPs, a far greater proportion of IGF-1 LR3 remains free to engage the receptor. This is the molecular basis for IGF-1 LR3's dramatically enhanced potency in biological systems [1, 2, 3].
The practical consequence is that IGF-1 LR3 at nanomolar concentrations achieves the same biological effects that would require micromolar concentrations of native IGF-1 in IGFBP-containing environments such as serum-supplemented cell culture media or in vivo conditions.
Scientific Research Review
Foundational Studies: Francis et al. and the Development of LR3
The seminal work establishing IGF-1 LR3 as a viable research tool was published by Francis and colleagues at the Cooperative Research Centre for Tissue Growth and Repair in Adelaide, Australia. Their systematic studies characterized the binding properties and biological activity of the LR3 analog, demonstrating that the N-terminal extension and Arg3 substitution together reduced IGFBP binding by approximately two orders of magnitude without compromising IGF-1R activation [2].
Key findings from this foundational work included:
Binding specificity: IGF-1 LR3 showed less than 1% of native IGF-1's affinity for IGFBP-1, IGFBP-2, IGFBP-3, IGFBP-4, and IGFBP-5 in competitive binding assays, while retaining approximately 85-100% of native IGF-1's affinity for the IGF-1R.
Enhanced potency in cell proliferation: In cell culture systems containing serum (and therefore IGFBPs), IGF-1 LR3 was approximately 2-3 fold more potent than native IGF-1 in stimulating cell proliferation, consistent with its resistance to IGFBP sequestration.
Extended biological activity: In time-course experiments, IGF-1 LR3 maintained stimulatory activity for substantially longer periods than native IGF-1, reflecting its resistance to IGFBP-mediated clearance [2, 3].
Skeletal Muscle Hypertrophy Research
The IGF-1/IGF-1R signaling axis has been established as a critical regulator of skeletal muscle mass, and IGF-1 LR3 has been extensively used to study muscle hypertrophy mechanisms.
Satellite cell activation and proliferation: Skeletal muscle contains resident stem cells called satellite cells that are quiescent under normal conditions but activate in response to injury, exercise, or growth factor stimulation. IGF-1 signaling through the IGF-1R/PI3K/Akt pathway is a potent activator of satellite cell proliferation and differentiation into myogenic precursors. Studies using IGF-1 LR3 in primary satellite cell cultures have demonstrated dose-dependent increases in proliferation and subsequent fusion into multinucleated myotubes [16, 20].
Myotube hypertrophy: In differentiated C2C12 myotubes (an established muscle cell line), IGF-1 LR3 treatment activates mTORC1 signaling and increases protein synthesis rates, resulting in measurable increases in myotube diameter. This effect is blocked by the mTORC1 inhibitor rapamycin, confirming the dependence on mTOR-mediated protein synthesis [15, 21].
Muscle-specific IGF-1 overexpression studies: Transgenic mouse models overexpressing IGF-1 specifically in skeletal muscle (using the muscle creatine kinase promoter) develop significant muscle hypertrophy without the systemic metabolic complications of elevated circulating IGF-1. These models have demonstrated that local IGF-1 action in muscle is sufficient to drive hypertrophy through autocrine/paracrine mechanisms [22]. Although these studies used native IGF-1 transgenes, they established the biological rationale for using IGF-1 LR3 as a more potent exogenous stimulus for the same pathway.
Adams and Haddad (2001) demonstrated that mechanical loading of rat skeletal muscle increases local IGF-1 expression and that the resulting muscle hypertrophy depends on PI3K/Akt/mTOR signaling. This established that the IGF-1 pathway is activated by resistance exercise and mediates load-induced muscle growth [23].
Cell Culture and Bioprocessing Applications
One of the most practical applications of IGF-1 LR3 is as a cell culture media supplement, where its resistance to IGFBP sequestration provides significant advantages over native IGF-1.
Serum-free and reduced-serum culture: The biopharmaceutical industry's shift toward serum-free media for cell culture has created demand for defined growth factor supplements. IGF-1 LR3 is widely used in serum-free formulations for Chinese hamster ovary (CHO) cells, hybridomas, and other cell lines because it maintains cell viability and proliferation without the batch-to-batch variability and IGFBP interference associated with serum supplementation [24].
Extended media stability: Because IGF-1 LR3 resists IGFBP-mediated degradation and clearance, it maintains biological activity in conditioned media for longer periods than native IGF-1. This is particularly valuable in fed-batch and perfusion culture systems where growth factor stability directly affects process economics.
Stem cell maintenance: IGF-1 LR3 has been used in protocols for maintaining human embryonic stem cells (hESCs) and induced pluripotent stem cells (iPSCs) in undifferentiated states, as well as in directed differentiation protocols where IGF-1 signaling promotes specific lineage commitment [25].
Cancer Biology Research
The IGF-1R signaling pathway is one of the most studied axes in cancer biology, and IGF-1 LR3 has served as a valuable research tool for understanding how dysregulated IGF signaling contributes to tumorigenesis.
Mitogenic and anti-apoptotic effects: IGF-1R signaling promotes both cell proliferation (via MAPK) and survival (via PI3K/Akt), two hallmarks of cancer. Overexpression of IGF-1R and/or elevated IGF-1 levels have been epidemiologically associated with increased risk of breast, prostate, colorectal, and lung cancers [26].
Resistance to apoptosis: Cancer cells frequently exploit IGF-1R signaling to resist apoptosis induced by chemotherapeutic agents and radiation. Research using IGF-1 LR3 has demonstrated that IGF-1R activation can confer resistance to doxorubicin, cisplatin, and targeted therapies in multiple cancer cell lines, providing mechanistic insight into clinical drug resistance [27].
IGF-1R as a therapeutic target: The recognition of IGF-1R's role in cancer led to the development of anti-IGF-1R antibodies (such as figitumumab, dalotuzumab, and ganitumab) and IGF-1R tyrosine kinase inhibitors (such as linsitinib) as potential anticancer therapeutics. Studies using IGF-1 LR3 as a positive control agonist were instrumental in characterizing the pharmacology of these antagonists [28].
Wound Healing and Tissue Repair
IGF-1 is an established promoter of tissue repair, and IGF-1 LR3's enhanced potency has made it useful for studying wound healing mechanisms.
Fibroblast proliferation and collagen synthesis: IGF-1 LR3 stimulates dermal fibroblast proliferation and increases type I collagen synthesis through the PI3K/Akt pathway. These effects are relevant to wound closure and scar remodeling [29].
Chondrocyte metabolism: In cartilage biology, IGF-1 is the principal anabolic growth factor driving proteoglycan synthesis. IGF-1 LR3 has been used in ex vivo cartilage explant cultures to study the regulation of matrix synthesis and degradation, with applications in osteoarthritis research [30].
Bone regeneration: IGF-1 signaling promotes osteoblast proliferation and differentiation, and suppresses osteoclast activity. Research with IGF-1 analogs including LR3 has explored potential applications in fracture healing and bone tissue engineering [31].
Metabolic Research
The insulin-like metabolic activity of IGF-1 makes it relevant to diabetes and metabolic syndrome research.
Glucose uptake: IGF-1 LR3, like native IGF-1, can stimulate glucose uptake in skeletal muscle cells through a mechanism involving GLUT4 translocation. Although IGF-1R activation produces this effect primarily through the PI3K/Akt pathway (similar to insulin receptor signaling), IGF-1 LR3 can also cross-react weakly with the insulin receptor at supraphysiological concentrations [32].
Insulin sensitization: Studies in insulin-resistant cell models have shown that IGF-1R activation can partially compensate for impaired insulin receptor signaling, improving glucose metabolism through convergent downstream pathways [33].
Comparison with Related Growth Factors
IGF-1 LR3 vs. Native IGF-1
| Parameter | Native IGF-1 | IGF-1 LR3 |
|---|---|---|
| Amino acids | 70 | 83 |
| Molecular weight | Approximately 7,649 Da | Approximately 9,111 Da |
| IGFBP binding affinity | High (Kd 0.1-10 nM) | Approximately 100-fold reduced |
| IGF-1R binding affinity | Kd approximately 0.2-1 nM | Near-native (approximately 85-100% retained) |
| Free peptide half-life | Approximately 12-15 minutes | Approximately 20-30 hours |
| Total half-life (IGFBP-bound) | Approximately 12-16 hours | N/A (minimal binding) |
| Potency in IGFBP-rich systems | Reduced by IGFBP sequestration | 2-3 fold greater than IGF-1 |
| Potency in IGFBP-free systems | Equivalent | Approximately equivalent |
| Primary use | Endocrine/clinical research | Cell culture, in vitro research |
IGF-1 LR3 vs. IGF-1 DES(1-3)
IGF-1 DES(1-3) is another IGF-1 analog designed to evade IGFBP binding, achieved by truncating the first three N-terminal amino acids (removing Gly-Pro-Glu):
| Parameter | IGF-1 LR3 | IGF-1 DES(1-3) |
|---|---|---|
| Amino acids | 83 | 67 |
| Modification strategy | N-terminal extension + Arg3 substitution | N-terminal truncation |
| IGFBP binding reduction | Approximately 100-fold | Approximately 100-fold |
| IGF-1R affinity | Near-native | Near-native |
| Stability | Good (extended half-life) | Moderate (shorter than LR3) |
| Cell culture utility | Widely used | Less commonly used |
| Mechanism of IGFBP evasion | Steric blockade + charge reversal | Removal of contact residues |
IGF-1 LR3 vs. GH-Releasing Peptides
Researchers studying anabolic signaling may encounter both IGF-1 LR3 and growth hormone-releasing peptides such as CJC-1295 and Ipamorelin. These operate through fundamentally different mechanisms:
| Parameter | IGF-1 LR3 | CJC-1295 / Ipamorelin |
|---|---|---|
| Target receptor | IGF-1R (direct) | GHRH-R / GHS-R (indirect) |
| Mechanism | Direct IGF-1R agonism | Stimulate GH release, then hepatic IGF-1 production |
| Pathway | IGF-1R → PI3K/Akt/mTOR | GHRH-R/GHS-R → GH → GHR → JAK2/STAT5 → IGF-1 gene |
| Onset of IGF-1R activation | Immediate | Delayed (requires GH secretion, then IGF-1 synthesis) |
| Specificity | IGF-1R only | Broad (GH has multiple IGF-1-independent effects) |
| IGF-1 elevation pattern | Exogenous, non-physiological | Pulsatile, physiological |
| Use case | Direct cell signaling studies | GH/IGF-1 axis research |
The key distinction is that CJC-1295 and Ipamorelin work upstream in the somatotropic axis by stimulating endogenous growth hormone secretion, which subsequently increases hepatic IGF-1 production. IGF-1 LR3 bypasses this entire axis, directly activating IGF-1R without requiring growth hormone signaling. This makes IGF-1 LR3 the preferred tool for studying IGF-1R-specific signaling, while GH secretagogues are better suited for studying the integrated GH-IGF-1 axis.
IGF-1 vs. IGF-2 and Insulin
| Parameter | IGF-1 LR3 | IGF-2 | Insulin |
|---|---|---|---|
| Amino acids | 83 | 67 | 51 (A+B chains) |
| Primary receptor | IGF-1R | IGF-1R, IGF-2R | Insulin receptor (IR) |
| PI3K/Akt activation | Strong | Moderate | Strong |
| MAPK activation | Strong | Moderate | Weak |
| Protein synthesis | Primary effect | Moderate effect | Moderate effect |
| Glucose uptake | Moderate | Weak | Primary effect |
| Mitogenic activity | High | High | Low |
| Anti-apoptotic activity | High | Moderate | Moderate |
Safety Profile and Pharmacology
Pharmacokinetic Properties
The pharmacokinetic profile of IGF-1 LR3 is defined by its engineered resistance to IGFBP binding:
Absorption: As a peptide of approximately 9.1 kDa, IGF-1 LR3 is not orally bioavailable and must be administered parenterally in research settings. Subcutaneous administration results in gradual absorption from the injection site into the systemic circulation.
Distribution: Unlike native IGF-1, which is approximately 99% bound to IGFBPs in circulation, IGF-1 LR3 circulates predominantly in the free form. This results in a much larger volume of distribution as the unbound peptide can readily distribute into tissues. The free form also means that IGF-1 LR3 has direct, immediate access to IGF-1R on target cells without requiring IGFBP proteolysis for release [3, 8].
Metabolism: IGF-1 LR3 is subject to proteolytic degradation by serum and tissue proteases. However, the absence of IGFBP-mediated clearance mechanisms and the peptide's resistance to IGFBP-directed proteases contribute to its extended functional half-life of approximately 20-30 hours [1].
Elimination: Renal filtration and hepatic proteolysis are the expected primary clearance mechanisms, consistent with other peptides of similar molecular weight.
Observed Effects in Preclinical Research
The following biological effects have been documented in preclinical research with IGF-1 or IGF-1 LR3 and should be considered in the context of research protocol design:
Hypoglycemia: IGF-1 possesses insulin-like metabolic activity and can suppress hepatic glucose production while stimulating peripheral glucose uptake. In research models, high doses of IGF-1 have produced clinically significant hypoglycemia. This effect is potentially more pronounced with IGF-1 LR3 due to its enhanced bioavailability [32, 33].
Organ growth: Sustained IGF-1R activation promotes proliferative and hypertrophic responses in multiple tissues. Chronic administration of IGF-1 in animal models has been associated with organomegaly, including cardiac, splenic, and renal enlargement [34].
Tumor promotion: Because IGF-1R signaling is a potent mitogenic and anti-apoptotic pathway, sustained activation could theoretically promote the growth of pre-existing neoplasms. Epidemiological studies have associated elevated circulating IGF-1 levels with modestly increased cancer risk in human populations [26]. This does not indicate that IGF-1 is carcinogenic (it does not initiate transformation), but rather that it may promote the survival and proliferation of cells that have already undergone malignant transformation.
Electrolyte shifts: IGF-1 administration has been associated with fluid retention and alterations in electrolyte balance, including phosphate redistribution, in clinical research [34].
Considerations for Research Protocol Design
Researchers using IGF-1 LR3 should be aware of several factors relevant to experimental design:
Concentration-response relationships: Due to its IGFBP resistance, the effective concentration of IGF-1 LR3 differs substantially from native IGF-1 in systems containing IGFBPs. Dose-response curves should be established empirically for each experimental system rather than extrapolated from native IGF-1 literature.
Serum interactions: In serum-supplemented media, endogenous IGFBPs from serum will partially sequester native IGF-1 but have minimal effect on IGF-1 LR3. This must be considered when comparing the two molecules experimentally.
Cross-reactivity: At high concentrations, IGF-1 LR3 can weakly activate the insulin receptor (IR), particularly the IR-A isoform. Researchers studying IGF-1R-specific signaling should use concentrations within the selective range (typically less than 100 ng/mL) or include insulin receptor-specific controls [13].
Stability: IGF-1 LR3 should be reconstituted in acidic buffer (pH 3-4) or at physiological pH in the presence of carrier protein (e.g., 0.1% BSA) to prevent adsorption to plastic surfaces and maintain stability. Repeated freeze-thaw cycles should be avoided.
Research Applications
Cell Biology and Culture Applications
IGF-1 LR3 has become a standard component in modern cell culture formulations, particularly in the following contexts:
1. Serum-free media supplementation: The biopharmaceutical industry's adoption of serum-free and chemically defined media for therapeutic protein production has created widespread demand for IGF-1 LR3 as a growth factor supplement. It provides IGF-1R-mediated survival and proliferative signals without the variability and contamination risks of serum. Typical working concentrations range from 20-100 ng/mL in serum-free formulations [24].
2. Stem cell culture and differentiation: IGF-1 LR3 is used in protocols for maintaining pluripotent stem cells and directing their differentiation toward specific lineages, including myogenic, osteogenic, and neural fates. The PI3K/Akt pathway activation provided by IGF-1R signaling is a key component of many differentiation cocktails [25].
3. Primary cell culture: Primary cells harvested from tissues often require growth factor support to survive ex vivo. IGF-1 LR3 is particularly useful for primary satellite cells, fibroblasts, chondrocytes, and osteoblasts, where it promotes both survival and proliferation.
4. Hybridoma culture: Antibody-producing hybridoma cells used for monoclonal antibody generation benefit from IGF-1 LR3 supplementation, which enhances cell viability and antibody production in reduced-serum conditions.
Muscle Biology Research
5. Myogenesis studies: IGF-1 LR3 is a standard tool for studying the molecular mechanisms of myogenesis, including satellite cell activation, myoblast proliferation, differentiation, and fusion. Its use in C2C12 cells and primary myoblast cultures has generated fundamental insights into the PI3K/Akt/mTOR pathway's role in muscle formation [16, 20, 21].
6. Hypertrophy signaling: Studies of muscle protein synthesis regulation use IGF-1 LR3 as a defined stimulus to activate mTORC1 and its downstream effectors S6K1 and 4E-BP1. These experiments have been critical for understanding how anabolic signals are translated into increased protein accretion [15].
7. Atrophy prevention research: IGF-1 LR3 has been used in models of muscle atrophy (disuse, denervation, cachexia) to study whether activation of anabolic IGF-1R signaling can counteract catabolic pathways involving the ubiquitin-proteasome system and autophagy-lysosome pathway [35].
Cancer Research
8. Oncogenic signaling studies: IGF-1 LR3 serves as a positive control agonist for studying IGF-1R-mediated signaling in cancer cell lines. Researchers use it to characterize how IGF-1R activation drives proliferation, survival, migration, and invasion in various tumor types [26, 27].
9. Drug resistance research: The role of IGF-1R signaling in conferring resistance to chemotherapy, radiation, and targeted therapies is studied using IGF-1 LR3 to activate the pathway in the presence of cytotoxic agents [28].
10. Anti-IGF-1R drug development: Pharmaceutical screening programs targeting IGF-1R use IGF-1 LR3 as the reference agonist for characterizing receptor antagonists, kinase inhibitors, and antibody therapeutics.
Metabolic and Endocrine Research
11. Insulin-IGF-1 cross-talk: The overlapping signaling of insulin and IGF-1 pathways is studied using IGF-1 LR3 alongside insulin to dissect receptor-specific versus shared downstream effects on glucose metabolism, lipid synthesis, and gene expression [32, 33].
12. GH-IGF-1 axis research: While IGF-1 LR3 bypasses the GH axis, it is used as a comparative tool alongside GH secretagogues such as CJC-1295 and Ipamorelin to distinguish between GH-dependent and IGF-1-dependent effects in growth and metabolism research.
Tissue Engineering and Regenerative Medicine
13. Scaffold-based tissue engineering: IGF-1 LR3 is incorporated into biomaterial scaffolds (hydrogels, electrospun fibers, 3D-printed matrices) to provide sustained growth factor delivery for tissue engineering applications. Its stability and potency make it well-suited for controlled-release formulations [31].
14. Cartilage repair models: In vitro and ex vivo cartilage tissue engineering studies use IGF-1 LR3 to stimulate chondrocyte proliferation and extracellular matrix synthesis, with relevance to osteoarthritis and cartilage defect repair [30].
References
[1] Francis, G.L., Ross, M., Ballard, F.J., Milner, S.J., Senn, C., McNeil, K.A., Wallace, J.C., King, R., & Wells, J.R. (1992). "Novel recombinant fusion protein analogues of insulin-like growth factor (IGF)-I indicate the relative importance of IGF-binding protein and receptor binding for enhanced biological potency." Journal of Molecular Endocrinology, 8(3), 213-223. DOI: 10.1677/jme.0.0080213
[2] Francis, G.L., Aplin, S.E., Milner, S.J., McNeil, K.A., Ballard, F.J., & Wallace, J.C. (1993). "Insulin-like growth factor (IGF)-II binding to IGF-binding proteins and IGF receptors is modified by deletion of the N-terminal hexapeptide or substitution of arginine for glutamate-3 in IGF-I." Biochemical Journal, 293(Pt 3), 713-719. DOI: 10.1042/bj2930713
[3] Tomas, F.M., Knowles, S.E., Owens, P.C., Chandler, C.S., Francis, G.L., Read, L.C., & Ballard, F.J. (1993). "Insulin-like growth factor-I (IGF-I) and especially IGF-I variants are anabolic in dexamethasone-treated rats." Biochemical Journal, 292(Pt 3), 781-786. DOI: 10.1042/bj2920781
[4] Ballard, F.J., Wallace, J.C., Francis, G.L., Read, L.C., & Tomas, F.M. (1996). "Des(1-3)IGF-I: a truncated form of insulin-like growth factor-I." International Journal of Biochemistry & Cell Biology, 28(10), 1085-1087. DOI: 10.1016/1357-2725(96)00056-8
[5] Salmon, W.D. Jr. & Daughaday, W.H. (1957). "A hormonally controlled serum factor which stimulates sulfate incorporation by cartilage in vitro." Journal of Laboratory and Clinical Medicine, 49(6), 825-836. DOI: 10.5555/uri:pii:0022214357900504
[6] Rinderknecht, E. & Humbel, R.E. (1978). "The amino acid sequence of human insulin-like growth factor I and its structural homology with proinsulin." Journal of Biological Chemistry, 253(8), 2769-2776. DOI: 10.1016/S0021-9258(17)40889-1
[7] Laron, Z. (2004). "Laron syndrome (primary growth hormone resistance or insensitivity): the personal experience 1958-2003." Journal of Clinical Endocrinology & Metabolism, 89(3), 1031-1044. DOI: 10.1210/jc.2003-031033
[8] Jones, J.I. & Clemmons, D.R. (1995). "Insulin-like growth factors and their binding proteins: biological actions." Endocrine Reviews, 16(1), 3-34. DOI: 10.1210/edrv-16-1-3
[9] Rajaram, S., Baylink, D.J., & Mohan, S. (1997). "Insulin-like growth factor-binding proteins in serum and other biological fluids: regulation and functions." Endocrine Reviews, 18(6), 801-831. DOI: 10.1210/edrv.18.6.0321
[10] Clemmons, D.R. (2001). "Use of mutagenesis to probe IGF-binding protein structure/function relationships." Endocrine Reviews, 22(6), 800-817. DOI: 10.1210/edrv.22.6.0449
[11] De Meyts, P. & Whittaker, J. (2002). "Structural biology of insulin and IGF1 receptors: implications for drug design." Nature Reviews Drug Discovery, 1(10), 769-783. DOI: 10.1038/nrd917
[12] Sato, A., Nishimura, S., Ohkubo, T., Kyogoku, Y., Koyama, S., Kobayashi, M., Yasuda, T., & Kobayashi, Y. (1993). "Three-dimensional structure of human insulin-like growth factor-I (IGF-I) determined by 1H-NMR and distance geometry." International Journal of Peptide and Protein Research, 41(5), 433-440. DOI: 10.1111/j.1399-3011.1993.tb00462.x
[13] Adams, T.E., Epa, V.C., Garrett, T.P., & Ward, C.W. (2000). "Structure and function of the type 1 insulin-like growth factor receptor." Cellular and Molecular Life Sciences, 57(7), 1050-1093. DOI: 10.1007/PL00000744
[14] Datta, S.R., Brunet, A., & Greenberg, M.E. (1999). "Cellular survival: a play in three Akts." Genes & Development, 13(22), 2905-2927. DOI: 10.1101/gad.13.22.2905
[15] Rommel, C., Bodine, S.C., Clarke, B.A., Rossman, R., Nunez, L., Stitt, T.N., Yancopoulos, G.D., & Glass, D.J. (2001). "Mediation of IGF-1-induced skeletal myotube hypertrophy by PI(3)K/Akt/mTOR and PI(3)K/Akt/GSK3 pathways." Nature Cell Biology, 3(11), 1009-1013. DOI: 10.1038/ncb1101-1009
[16] Coolican, S.A., Samuel, D.S., Ewton, D.Z., McWade, F.J., & Bhatt, H.M. (1997). "The mitogenic and myogenic actions of insulin-like growth factors utilize distinct signaling pathways." Journal of Biological Chemistry, 272(10), 6653-6662. DOI: 10.1074/jbc.272.10.6653
[17] Brunet, A., Bonni, A., Zigmond, M.J., Lin, M.Z., Juo, P., Hu, L.S., Anderson, M.J., Arden, K.C., Blenis, J., & Greenberg, M.E. (1999). "Akt promotes cell survival by phosphorylating and inhibiting a Forkhead transcription factor." Cell, 96(6), 857-868. DOI: 10.1016/S0092-8674(00)80595-4
[18] Skolnik, E.Y., Batzer, A., Li, N., Lee, C.H., Lowenstein, E., Mohammadi, M., Margolis, B., & Schlessinger, J. (1993). "The function of GRB2 in linking the insulin receptor to Ras signaling pathways." Science, 260(5116), 1953-1955. DOI: 10.1126/science.8316835
[19] Chitnis, M.M., Yuen, J.S., Protheroe, A.S., Pollak, M., & Macaulay, V.M. (2008). "The type 1 insulin-like growth factor receptor pathway." Clinical Cancer Research, 14(20), 6364-6370. DOI: 10.1158/1078-0432.CCR-07-4879
[20] Florini, J.R., Ewton, D.Z., & Coolican, S.A. (1996). "Growth hormone and the insulin-like growth factor system in myogenesis." Endocrine Reviews, 17(5), 481-517. DOI: 10.1210/edrv-17-5-481
[21] Bodine, S.C., Stitt, T.N., Gonzalez, M., Kline, W.O., Stover, G.L., Bauerlein, R., Zlotchenko, E., Scrimgeour, A., Lawrence, J.C., Glass, D.J., & Yancopoulos, G.D. (2001). "Akt/mTOR pathway is a crucial regulator of skeletal muscle hypertrophy and can prevent muscle atrophy in vivo." Nature Cell Biology, 3(11), 1014-1019. DOI: 10.1038/ncb1101-1014
[22] Musaro, A., McCullagh, K., Paul, A., Houghton, L., Dobrowolny, G., Molinaro, M., Barton, E.R., Sweeney, H.L., & Rosenthal, N. (2001). "Localized Igf-1 transgene expression sustains hypertrophy and regeneration in senescent skeletal muscle." Nature Genetics, 27(2), 195-200. DOI: 10.1038/84839
[23] Adams, G.R. & Haddad, F. (1996). "The relationships among IGF-1, DNA content, and protein accumulation during skeletal muscle hypertrophy." Journal of Applied Physiology, 81(6), 2509-2516. DOI: 10.1152/jappl.1996.81.6.2509
[24] Freshney, R.I. (2010). "Serum-free media." In Culture of Animal Cells: A Manual of Basic Technique and Specialized Applications (6th ed.). Wiley-Blackwell. DOI: 10.1002/9780470649367
[25] Bendall, S.C., Stewart, M.H., Menendez, P., George, D., Vijayaragavan, K., Werbowetski-Ogilvie, T., Ramos-Mejia, V., Rouleau, A., Yang, J., Bosse, M., Lajoie, G., & Bhutia, M. (2007). "IGF and FGF cooperatively establish the regulatory stem cell niche of pluripotent human cells in vitro." Nature, 448(7157), 1015-1021. DOI: 10.1038/nature06027
[26] Pollak, M.N., Schernhammer, E.S., & Hankinson, S.E. (2004). "Insulin-like growth factors and neoplasia." Nature Reviews Cancer, 4(7), 505-518. DOI: 10.1038/nrc1387
[27] Baserga, R. (2000). "The contradictions of the insulin-like growth factor 1 receptor." Oncogene, 19(49), 5574-5581. DOI: 10.1038/sj.onc.1203854
[28] Gualberto, A. & Pollak, M. (2009). "Emerging role of insulin-like growth factor receptor inhibitors in oncology: early clinical trial results and future directions." Oncogene, 28(34), 3009-3021. DOI: 10.1038/onc.2009.172
[29] Gartner, M.H., Benson, J.D., & Caldwell, M.D. (1992). "Insulin-like growth factors I and II expression in the healing wound." Journal of Surgical Research, 52(4), 389-394. DOI: 10.1016/0022-4804(92)90121-F
[30] Luyten, F.P., Hascall, V.C., Nissley, S.P., Morales, T.I., & Reddi, A.H. (1988). "Insulin-like growth factors maintain steady-state metabolism of proteoglycans in bovine articular cartilage explants." Archives of Biochemistry and Biophysics, 267(2), 416-425. DOI: 10.1016/0003-9861(88)90047-1
[31] Giustina, A., Mazziotti, G., & Canalis, E. (2008). "Growth hormone, insulin-like growth factors, and the skeleton." Endocrine Reviews, 29(5), 535-559. DOI: 10.1210/er.2007-0036
[32] Guler, H.P., Zapf, J., & Froesch, E.R. (1987). "Short-term metabolic effects of recombinant human insulin-like growth factor I in healthy adults." New England Journal of Medicine, 317(3), 137-140. DOI: 10.1056/NEJM198707163170303
[33] Moses, A.C., Young, S.C., Morrow, L.A., O'Brien, M., & Clemmons, D.R. (1996). "Recombinant human insulin-like growth factor I increases insulin sensitivity and improves glycemic control in type II diabetes." Diabetes, 45(1), 91-100. DOI: 10.2337/diab.45.1.91
[34] Jabri, N., Schalch, D.S., Schwartz, S.L., Fischer, J.S., Kipnes, M.S., Radnik, B.J., Turman, N.J., Marcsisin, V.S., & Guler, H.P. (1994). "Adverse effects of recombinant human insulin-like growth factor I in obese insulin-resistant type II diabetic patients." Diabetes, 43(3), 369-374. DOI: 10.2337/diab.43.3.369
[35] Sacheck, J.M., Ohtsuka, A., McLary, S.C., & Goldberg, A.L. (2004). "IGF-I stimulates muscle growth by suppressing protein breakdown and expression of atrophy-related ubiquitin ligases, atrogin-1 and MuRF1." American Journal of Physiology-Endocrinology and Metabolism, 287(4), E591-E601. DOI: 10.1152/ajpendo.00073.2004
Disclaimer
This article is for educational and informational purposes only. It is not intended as medical advice. IGF-1 LR3 is sold exclusively as a research peptide for in vitro and laboratory use. It is not intended for human consumption, therapeutic use, or as a dietary supplement. The information presented herein is derived from published peer-reviewed scientific literature and does not constitute medical recommendations. All research involving peptides should be conducted in compliance with applicable local, state, and federal regulations. Researchers should consult relevant institutional review boards and regulatory bodies before initiating any research protocols. Individual results may vary and past research findings do not guarantee future outcomes.
Published by BLL Peptides — Premium Research Peptides
IGF-1 LR3 is a research-grade long-acting Insulin-like Growth Factor 1 analog studied for potent anabolic effects on muscle protein synthesis, cellular proliferation, and glucose uptake via IGF-1 receptor signaling. Researchers investigating growth factor biology and anabolic physiology rely on pharmaceutical-grade purity for accurate data. Available at BLL Peptides — USA-made, rigorously tested.
| ✅ COA tested every batch | ✅ 98%+ purity guaranteed |
| ✅ USA manufactured, GMP-certified | ✅ Glass vials — not plastic |
| ✅ Veteran-owned company | ✅ Free shipping over $150 |